SpikeGLX Download Page
Synchronized acquisition from imec neural probes and NI-DAQ devices.
- Get compiled apps, support and help stuff on this page.
- See what's new on the site here.
- The source code repo is here.
This site supports prototype and currently shipping commercial products. The latest commercial PXI-based components are:
- Hardware:
Neuropixels 1.0, 2.0, NHP, UHD, Quad-base.- Software:
SpikeGLX API4.
What Is SpikeGLX?
SpikeGLX is a recording system for extracellular neural probes. The emphasis is on concurrent synchronous recording from high channel count probes together with numerous auxiliary analog and digital channels:
- Concurrent, synchronized acquisition from imec probes, OneBoxes and NI-DAQ devices.
- Supports all Neuropixels probe generations and types.
- Supports HHMI/Whisper system.
- Integration with live anatomical tracking: Pinpoint; Trajectory Explorer.
- Flexible visualization, filtering and sorting tools.
- Programmable triggering.
- Remote control and extensibility via MATLAB, C++, C, C#, Python.
- Powerful offline viewing and editing.
In addition to the SpikeGLX acquisition application, we provide a large complement of post-processing tools that are also found on this page. Our tool set is maintained and updated in lock step with SpikeGLX to make sure the tools work together correctly and have full awareness of all the flexibility that SpikeGLX provides in selecting parameters and making custom output files.
Note: If you want to craft your own post-processing tools, we strongly advise that you incorporate or at least consult the metadata parsing methods we have provided for both Python and MATLAB: SpikeGLX_Datafile_Tools.
System Requirements
(What to Buy)
Requirements differ according to platform:
OneBox Notes: Imec began selling OneBoxes October 5, 2024. Get data sheets and submit purchase requests at the Neuropixels site. OneBox does not depend upon any PXI hardware, hence, it does not require enclustra drivers.
Important!: The enclustra drivers required for PXI do not work with AMD-based computers. The PXI-based imec hardware can only run on computers using Intel CPUs and chipsets.
Important!: Older imec modules (shipped before Q2 2023) are programmed to work only in X4 and higher chassis slots. Learn more about this issue on the site's Firmware tab.
Latest Application Downloads
SpikeGLX API4 (PXI, OneBox, all probe types later than 3A):
Latest release highlights:
- Replaces all previous releases.
- Supports OneBox.
- Supports all probes including Quadbase.
- Does whole probe activity surveys.
- Has newer graphical site selection tools.
- Supports the (now separate) {MATLAB, C++, C, C#, Python} remote SDKs.
- Has many GUI enhancements and better help.
Important: Please read the
READMEfile in your download.
- Release 20251218-api4...Readme : Export survey, SDK selectDevices, api v4.1.3
Release downloads come with two executables:
- SpikeGLX.exe: Runs {imec probes, OneBox, NI, FileViewer}, but you need to install NI drivers to run it.
- SpikeGLX_NISIM.exe: Runs {imec probes, OneBox, FileViewer} but not NI hardware. Does not need NI drivers. Use this on your laptop to review data and plan new runs.
Suggested organization: Create folder 'SpikeGLX' on your desktop or
C:\then download/unzip associated stuff into it. It's fine to have several versions there as long as you run one at a time:
SpikeGLX\
Release_v20251218-api4.zip
Release_v20250325-phase30.zip
Release_v20251218-api4\
Release_v20250325-phase30\
Drivers\
Enclustra_Windows_10&11\
Tools\
Etc.\
PXI Enclustra Drivers
These required driver files are not included in your SpikeGLX release.
- Click the link below for your Windows OS version.
- Unzip the folder into your local folder of SpikeGLX-related stuff.
- Follow the installation ReadMe in the download.
Terminology: 'Enclustra' is a company that markets other vendor's FPGAs along with development and support tools. There are several FPGAs used in the Neuropixels hardware, including a Xilinx Zynq model, often referred to as 'the enclustra'.
Important!: The enclustra drivers required for PXI do not work with AMD-based computers. The PXI-based imec hardware can only run on computers using Intel CPUs and chipsets.
Remote Control/Scripting SDK
Note:
SDK = Software Development Kit
API = Application Programming Interface
SpikeGLX can be driven in real time by your own remote Windows or Linux application. The API uses the TCP/IP protocol. Your custom application can run on a separate computer (on the same network as SpikeGLX) or on the same computer. The API allows you to:
- Set/get parameters.
- Start/stop runs.
- Fetch data with low latency (<4 ms on same computer).
- Do many things you can do from the SpikeGLX GUI.
There are two SDKs:
- SpikeGLX-MATLAB-SDK : For MATLAB programs.
- SpikeGLX-CPP-SDK : For C++, C, C#, Python.
Get these at the source code repo here.
HelloSGLX (Windows)
The above SDKs are full featured and support developers writing in MATLAB, C++, C, C# and Python. If you are writing in anything else, you can use HelloSGLX instead. This is a command-line application that stands in between your script and SpikeGLX. For each command or query, your script uses its shell commands to launch HelloSGLX as a separate process. This is too slow for real-time data interaction, but still supports easy integration of SpikeGLX into an automated workflow. You can get/set run parameters and start/stop runs and file writing. Get it here.
Note that all of these SDKs require SpikeGLX 20220101 or later.
Command-line Tool Installation
- CatGT
- TPrime
- NIScaler
- C_Waves
- OverStrike
These come as zip files. To install one:
- Extract the contained folder to any destination you like.
- Keep the folder contents together, as you do with the SpikeGLX folder.
- Read the contained
ReadMe.txtfor instructions. - Look at the
runit.batexample batch script.
CatGT
CatGT is a command-line tool that does the following offline operations on SpikeGLX output files (all probe types):
- Optionally join trials with given run_name and index ranges [ga,gb] [ta,tb]...
- ...Or run on any individual file.
- Optionally apply demultiplexing corrections.
- Optionally apply band-pass and CAR filters.
- Optionally edit out saturation artifacts.
- By default extract tables of sync waveform edge times to drive TPrime.
- Optionally extract tables of other nonneural event times to be aligned with spikes.
- Optionally join the above outputs across different runs (supercat feature).
- CatGT: Tshift, CAR, Gfix
TPrime
TPrime is a command-line tool that maps event times (all probe types):
- Map time from one SpikeGLX data stream to any other.
- Translate all events to a single common timeline.
- Uses sync edges to achieve 1 to 2 sample accuracy.
- Maps TTL events extracted using CatGT.
- Maps spike times from any sorter.
- Sync: Aligning with Edges.
Post-processing Tools
SpikeGLX_Linux_offline is an AppImage that lets you run the offline FileViewer and ShankViewer natively under Linux.
NIScaler command-line tool to calibrate/rescale NI analog voltages acquired with SpikeGLX versions earlier than 20220101. SpikeGLX versions 20220101 and later automatically acquire scaled data.
Note: Only the Windows NIScaler tool can scan NI hardware and create calibration files. Both tools can apply calibration files to existing runs.
Measured correction size: {6133=0%, 6363=5%, 6341=7%}.
MATLAB and Python tools for parsing SpikeGLX meta and binary datafiles (supports all probes except quad-base).
MATLAB and Python tools that augment SpikeGLX metadata:
- Generates channel map files needed by Kilosort and JRClust (all SpikeGLX versions).
- Makes SpikeGLX data compatible with SpikeInterface (needed for versions prior to 20230326).
- Works for all probe types.
Average cluster waveforms and statistics command-line tool. This can be run separately like CatGT or used with ecephys_spike_sorting.
Jennifer Colonell's version of the Allen Institute ecephys_spike_sorting pipeline. This Python script-driven pipeline chains together: CatGT, KS2, Noise Cluster Tagging, C_Waves, QC metrics, TPrime.
Overwrite nasty parts of your binary data with zeros or lines using OverStrike. This helps remove artifacts that are too tricky for CatGT.
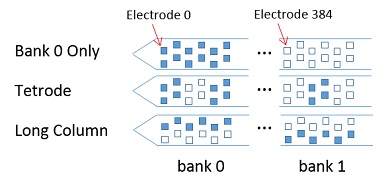
Interesting Map Files
- Tetrodes Bank Selection 3B (tetrodes covering banks zero and one)
- Long Column Bank Selection 3B (one column through banks zero and one)
Related help:
Metadata Guides
Descriptions of metafile items for each phase:
Questions or Problems?
1) For general How-To questions please consult the help documents and videos on this site, or pose your question to the Neuropixels Discord server so everyone can learn together. If there is any problem with the Discord invitation, email the administrator.
2) If the community can't help, try emailing me directly. I'm also happy to hear about new feature requests, and very eager to hear about suspected bugs. This is the best way to let me know there's a possible issue without causing alarm in the whole community. You can get my email from the About Box in the SpikeGLX application.
3) As a last resort you can register your observations of serious bugs/mistakes/errors using the SpikeGLX issue list on GitHub.
Help
- Neuropixels Discord server
- Neuropixels Support Page
- Noise: Reference and Ground
- Noise: Learn How To Solder
- Sharpening Apparatus (MS.Word.docx)
- PXIe Quickstart
- OneBox Quickstart
- Quadbase Quickstart
- SpikeGLX Tutorials
- SpikeGLX UserManual
- SpikeGLX Dirs and Files
- SpikeGLX Broken Shank Support
- SpikeGLX QuickRef
- SpikeGLX UCL Course 2025
- SpikeGLX UCL Course 2024
- SpikeGLX FAQ
- Metadata Help
- CatGT ReadMe
- CatGT: Tshift, CAR, Gfix
- Sync: Aligning with Edges
- IMRO Table Anatomy
How-to Videos
- Handle & Solder
- Sharpen Probe
- Install Software
- First Run
- Sync
- Graphical Probe IMRO Editor
- Whole-Probe Activity Survey
Licensing
Use is subject to Janelia Research Campus Software Copyright terms: http://license.janelia.org/license.
QLed components are subject to GNU Library GPL v2.0 terms, which are described here: https://github.com/billkarsh/SpikeGLX/blob/master/QLed-LGPLv2-LICENSE.txt.
fin